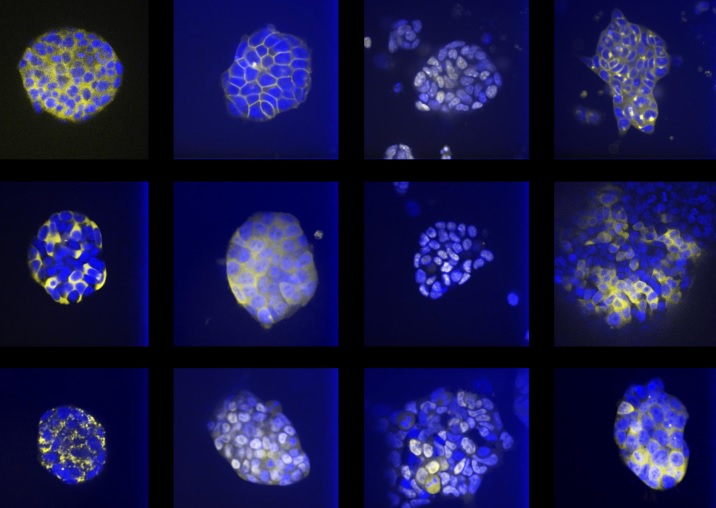
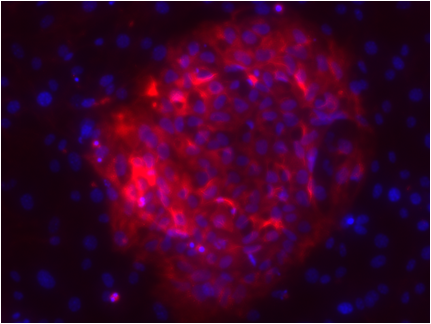
In our ERC project, we aim to understand chromatin plasticity and the function of non-polyadenylated transcription in pluripotency. We are combining biochemistry, single cell advanced imaging assays and high throughput technologies to study chromatin and transcription at a genome-wide scale in ES cells and during differentiation and reprogramming, as well as decipher the function of candidate chromatin proteins and candidate non-polyadenylated transcripts in pluripotency.

Collaborators:
Gil Ast, Tel-Aviv University
Gustavo Mostoslavsky, Boston University
Karsten Rippe, DKFZ
Paola Scaffidi, London Research Institute
Newman Sze, Nanyang Technological University, Singapore
Takumi Takizawa, Gunma University, Japan